You probably know the difference between in vitro and in vivo research. The in vitro model is laboratory-based; experiments occur in test tubes or Petri dishes. The in vivo model, in contrast, uses living organisms (animals or humans) to test the actions of molecules or molecular sets. But are you familiar with a third experimental model currently playing an important role in SARS-CoV-2 research?
What is an in silico study?
In silico is a modern technique. The pseudo-Latin term first appeared in 1987, taking its place among other better-known experimental terms: in vitro, in vivo and in situ. As you may have guessed, the silico part refers to silicon chips in computers. This technique uses all the power of modern bioinformatics tools: databases, quantitative structure-activity relationships, molecular modeling approaches, and machine learning. It is often an initial step in pharmaceutical development, and the information gained depends on the sophistication of the programs used, as well as the expertise of the team using them. What makes this model interesting is that, by using computers, it allows you to quickly test possibilities to see, conceptually, what works.
However, the in silico model has an important limitation – what happens in a computer model is not necessarily representative of what will happen in vitro and especially in vivo. An in silico result therefore cannot constitute a definitive prediction of the behavior of a molecule within the organism. It is an initial step in the long chain of scientific validation necessary to understand the effects of a molecule on a living being.
In silico work can be a cost effective and time efficient way of identifying molecules of interest, in order to plan in vitro experiments. It can also show that existing pharmaceuticals may be appropriate for a new purpose. In many cases, medications developed for one disease have been repurposed for another after an in silico study (Gordon et al. 2020; Kalathiya et al. 2020). A score is assigned to the matches called the “binding affinity”, and a higher score means that molecule binds better than a lower scored molecule. A higher affinity score in the in silico study means the molecule is more likely to bind in in vitro studies, and demonstrate the expected action in in vivo studies.
In the context of the SARS-CoV-2/COVID-19 pandemic, in silico studies have been key to quickly identify existing medications to use with COVID-19 patients. Likewise, in silico studies examining aromatic compounds could identify those effective against infection with this virus.
How does the SARS-CoV-2 virus infect humans?
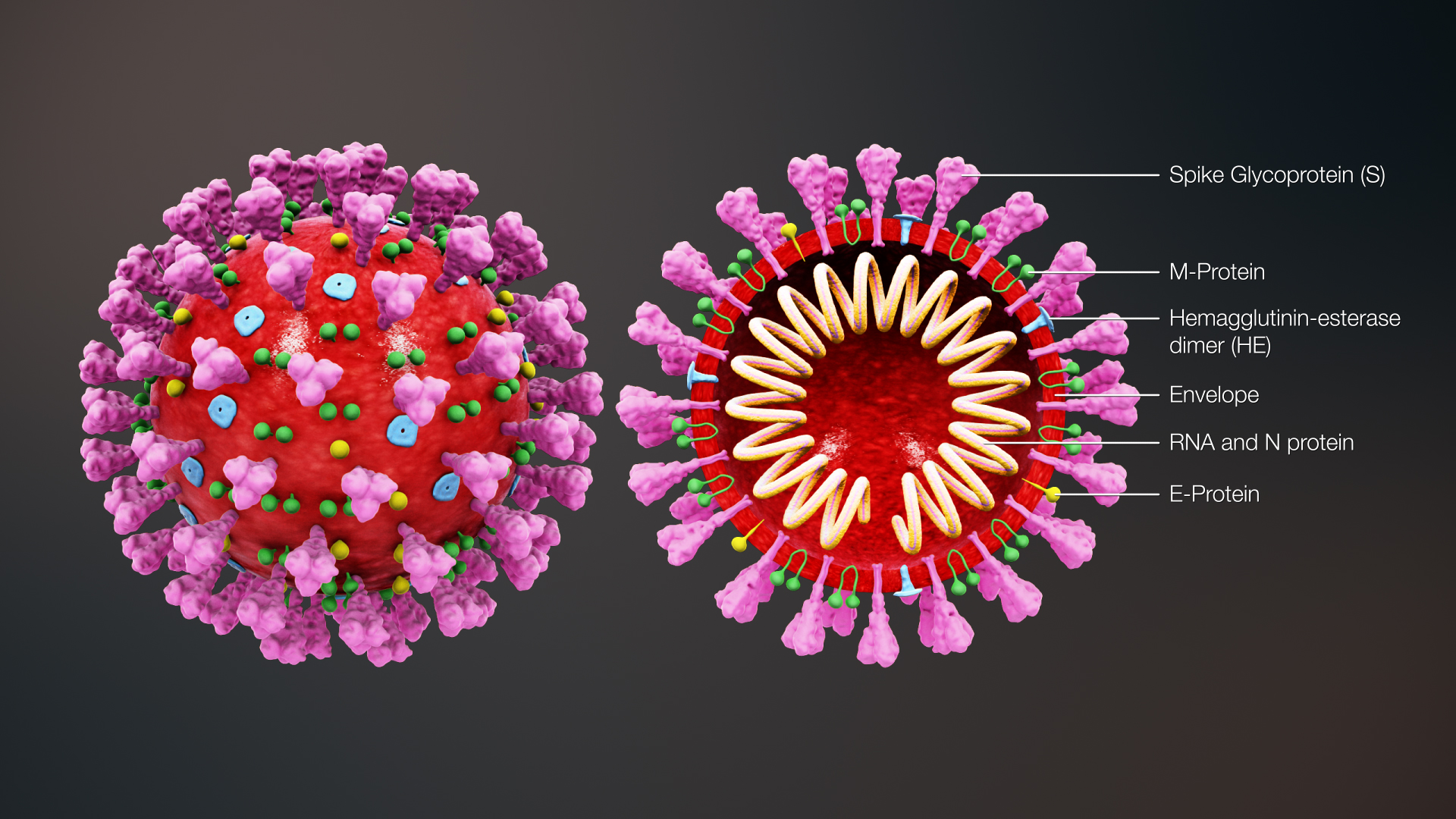
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has membrane proteins called spike proteins (S proteins) that are shaped like crowns/spikes. SARS-CoV-2 S proteins bind host cell angiotensin-converting enzyme 2 (ACE2) receptors (Hoffmann et al. 2020), which are notably found in alveolar epithelial cells of the lungs, but also in many other tissues including the small intestine, kidneys, heart, colon, and liver (Dehelean et al. 2020; Li et al. 2020). This means that all these tissues are potential targets for viral infections.
ACE2 binding is dependent on the S1 subunit of the S protein, which contains the receptor binding domain (RBD) (Hoffmann et al. 2020). The S2 subunit is cleaved from the larger protein by the host cell’s membrane-bound protease, called transmembrane protease serine 2 (TMPRSS2) (Hoffmann et al. 2020; Mollica et al. 2020). S1/S2 cleavage must occur for the viral membrane to fuse to the cell’s membrane, which is a necessary step for the virus to enter the host cell (Hoffmann et al. 2020; Mollica et al. 2020). These two enzymes (ACE2 and TMPRSS2) may be targets for COVID-19 treatments (Trezza et al. 2020), however, inhibiting host proteases could have adverse consequences (Tiwari et al. 2020).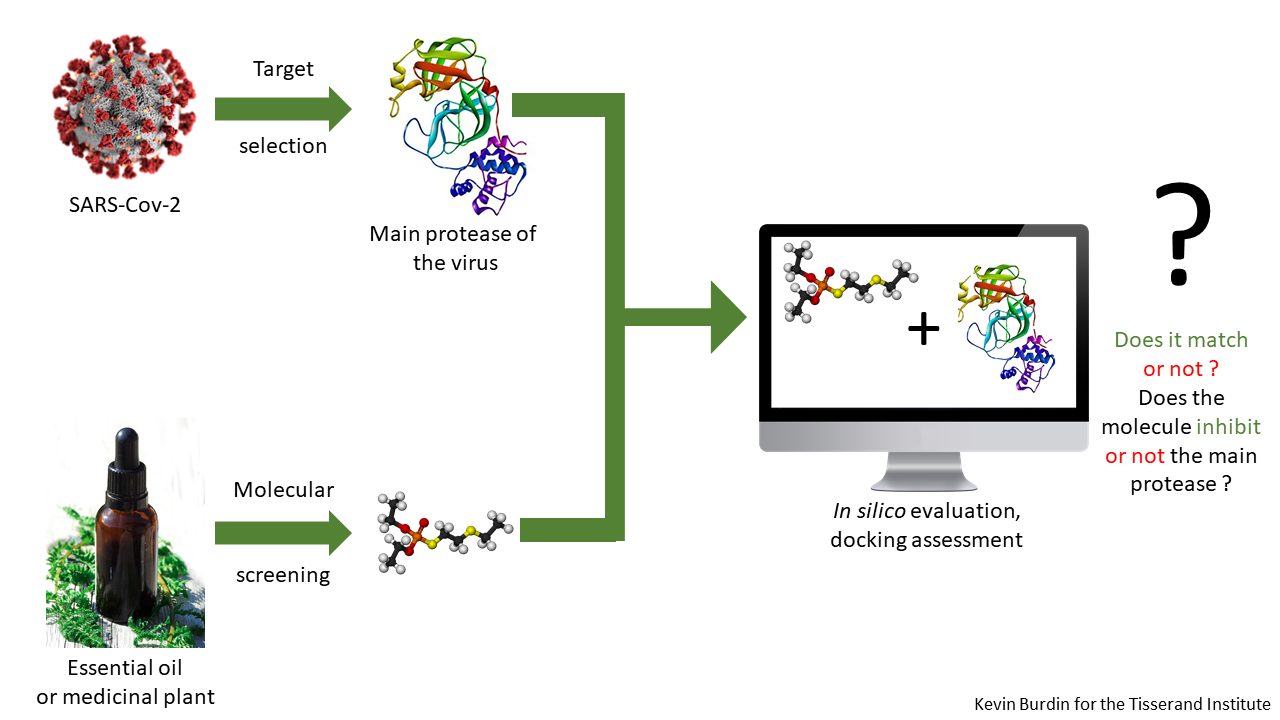
Upon entry, the virus is uncoated and its RNA is reproduced using the host cell’s cellular machinery. A viral enzyme called the coronavirus chymotrypsin-like protease or main protease (3CLpro or Mpro) is encoded in the viral genome, and is highly conserved in the beta coronaviruses (like MERS and SARS) (Qamar et al. 2020). 3CLpro is involved in the regulation of viral RNA replication (Yang et al. 2005), and 3CLpro inhibitors are being developed as COVID-19 treatments (Jin et al. 2020). New viral particles incorporate the host cell’s membrane as their own upon exit from the host cell, and these new viral particles are able to infect new cells. For more details, see previous review here.
Below are summaries of some of the recent in silico investigations of essential oil constituents or whole essential oils and their potential effect on each of the stages of COVID-19.
ACE2
An in silico study examined ACE2 inhibition by compounds in the distilled oil of Ammoides verticillata, a plant found in Algeria, and FDA approved drugs Captopril and Chloroquine (Abdelli et al. 2020). The essential oil is carvacrol-rich (51.2%), and also contains p-cymene (14.1%), thymol (13.0%), limonene (11.9%) and γ-terpinene (6.8%), but it is not commercially produced. Docking models of these compounds suggest that carvacrol is an ACE2 inhibitor comparable to Captopril and Chloroquine (Abdelli et al. 2020). Additional studies examining commercially available carvacrol-rich essential oils, like oregano (Origanum vulgare) could clarify whether carvacrol-rich essential oils can be used as ACE2 inhibitors.
Coronavirus chymotrypsin-like protease/main protease (3CLpro or Mpro)
In silico screens of chemical compound libraries to identify those that bind to coronavirus 3CLpro, and would thus inhibit viral replication, have been done (Yang et al. 2005; Jin et al. 2020).
Sharma and Kaur investigated 1,8-cineole, also known as eucalyptol, by conducting a molecular docking study using 3CLpro structures (Sharma and Kaur 2020a). 1,8-cineole is known for its antiviral properties and as an expectorant of bronchial secretions (Astani et al. 2010; Li et al. 2016; Yadav and Chandra 2017). The object of this study was to calculate the inhibition exerted by this molecule on 3CLpro. The study concludes 1,8-cineole may inhibit the viral protease, thereby inhibiting RNA replication. However, this study was published online by the authors before undergoing peer review, and has not been published in any peer reviewed journals. It is unclear whether their conclusions are significant. In addition, the same group published a brief online without peer review that erroneously claimed a compound called jensenone is found in “eucalyptus oil” (Sharma and Kaur 2020b). This compound is not found in Eucalyptus globulus essential oil (Tisserand and Young 2014), and has only been found in the distilled oil of an obscure plant called Melaleuca jensenii (Boland et al. 1992).
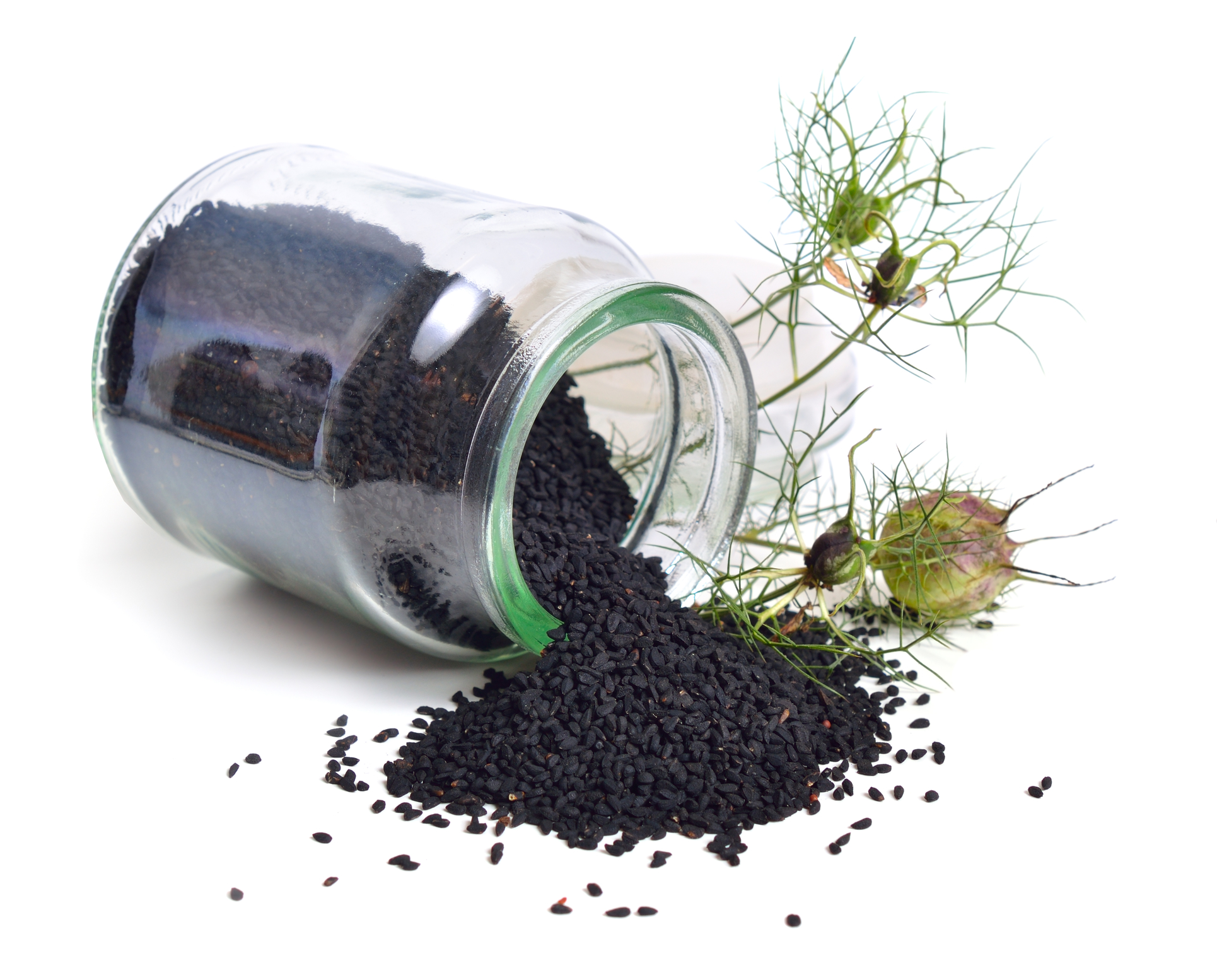
A study published without peer review, by Bouchentouf and Missoum, examined the effects of both volatile and non-volatile compounds of Nigella sativa (Black cumin, or Black seed) on SARS-CoV-2 (Bouchentouf and Missoum 2020). The authors discuss the plant compounds without explaining the method of extraction, and because they discuss non-volatile components, it is likely that they are not examining an essential oil. The in silico study examined nigellicin, nigellidine, nigellimine, α-hederin, carvacrol, thymol, thymoquinone, and dithymoquinone and their interaction with the viral protease 3CLpro. The authors state that nigellidine and α-hederin are good candidates to inhibit the virus, with results similar to allopathic treatments under study. However, these compounds are not found in the essential oil, and without peer review, it is unclear whether their findings are significant. Further studies will be needed to validate any antiviral properties, and their therapeutic potential.
Multiple targets
Garlic (Allium sativum) essential oil containing allyl disulfide (28.4%), allyl trisulfide (22.8%), allyl (E)-1-propenyl disulfide (8.2%), allyl methyl trisulfide (6.7%), and diallyl tetrasulfide (6.5%) was examined in an in silico study to determine interactions with ACE2 and the PDB6LU7 protein (Thuy et al. 2020), which is part of the 3CLpro viral protease. This essential oil showed concentrations of chemical constituents that differed from previous studies, and this may be due to differences in growing conditions. Of note, the studied essential oil lacks diallyl disulfide and diallyl trisulfide, which are the major constituents of published garlic essential oil studies (Tisserand and Young 2014). Of the 18 identified compounds, 17 showed strong binding scores to ACE2 and the PDB6LU7 protein (Thuy et al. 2020), suggesting garlic essential oil is an ACE2 inhibitor and 3CLpro inhibitor. Further in vitro and clinical studies would need to be done to determine if the in silico results translate to a COVID-19 treatment. However, garlic essential oil is moderately toxic and is an anticoagulant (Tisserand and Young 2014).
My et al. (My et al. 2020) examined Vietnamese cajeput (Melaleuca cajuputi) essential oil’s ability to bind ACE2 and the PDB6LU7 protein (part of 3CLpro) in their in silico study. Of the 24 main constituents, ten inhibited both 3CLpro and ACE2. The four molecules with the highest activity were, in decreasing order of potency: α-terpineol, guaiol, linalool and 1,8-cineole. This is interesting, because guaiol is typically found in trace-1.2% concentration in cajeput essential oil (Tisserand and Young 2014), while the essential oil examined in this study contained 6.5% guaiol (My et al. 2020). The essential oil differs from the profile in Tisserand and Young (2014) in other ways as well. The studied essential oil contains 1,8-cineole (31.6%), α-terpineol (10.7%), β-eudesmol (6.8%), α-eudesmol (6.7%), guiaol (6.5%), γ-eudesmol (4.3%), bulnesol (1.9%), β-myrcene (0.9%), terpinen-4-ol (0.9%), and linalool (0.6%) (My et al. 2020). As such, It is unclear if the essential oil used in the study is representative of cajeput, or if it may be a different oil. Regardless, the compounds did have a significant binding score in the in silico study, and if an in vitro study examines a more typical cajeput essential oil, it could add significance to this finding.
An in silico study by da Silva et al. focused on 171 essential oil constituents to study their action on ACE2, RBD (ACE2 binding region of the viral S protein), and 3CLpro (Silva et al. 2020). This study also examined other viral targets: SARS-CoV-2 endoribonuclease, SARS-CoV-2 ADP-ribose-1”-phosphatase, and SARS-CoV-2 RNA-dependent RNA polymerase. The chemical components were chosen by reviewing the literature for antiviral action, and include many compounds known to have antiviral action against viruses such as influenza (several strains) and herpes simplex viruses (HSV-1 and HSV-2). The authors found that (E)-β-farnesene had the highest docking score to SARS-CoV-2 3CLpro, but also note that the scores for (E,E)-α-farnesene, (E,E)-farnesol, and (E)-nerolidol are good (Silva et al. 2020). The same compounds scored highly against SARS-CoV-2 endoribonuclease.
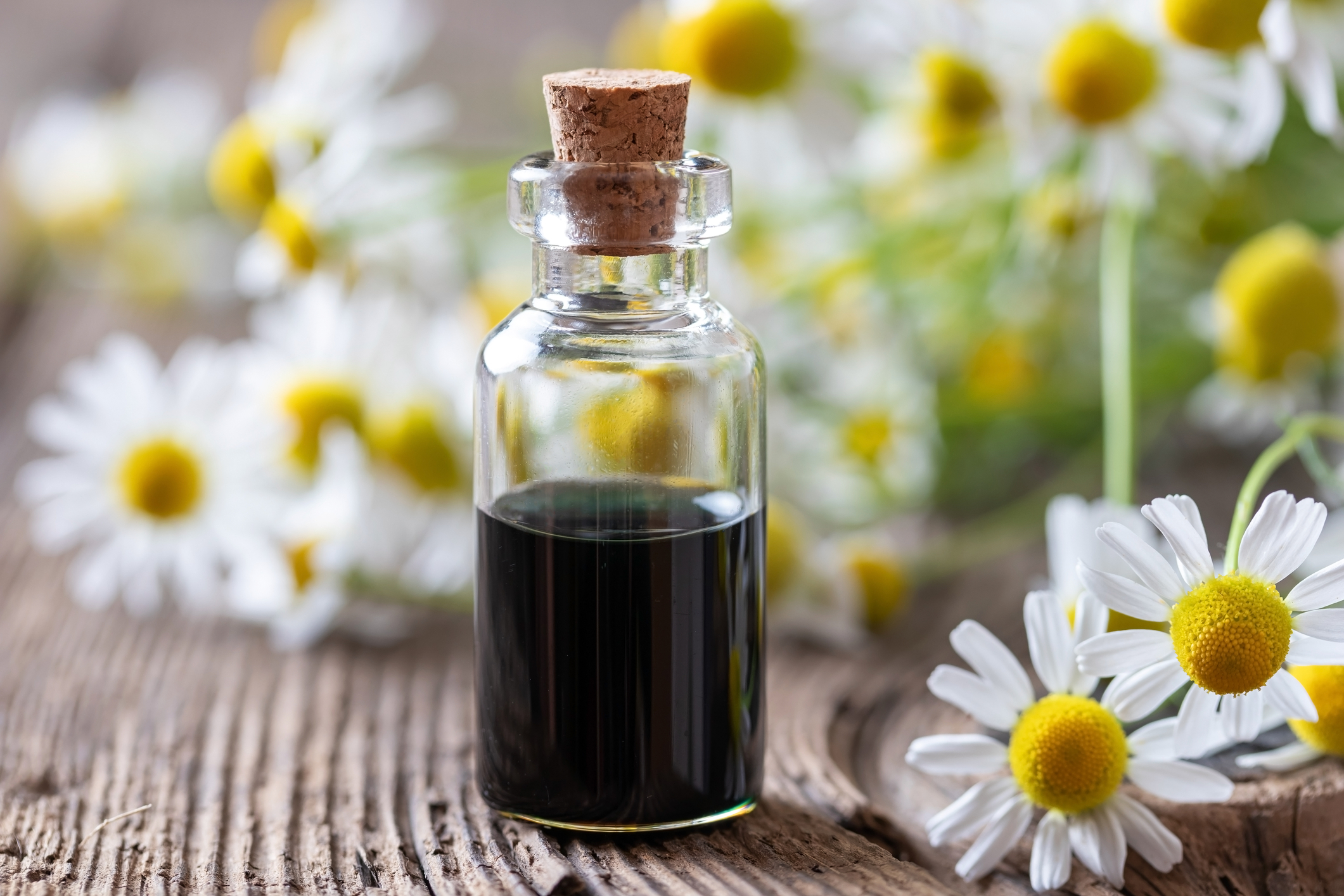
This is interesting, because the same compounds bind multiple targets in the study (Silva et al. 2020). German chamomile (Matricaria chamomilla) essential oil is typically β-farnesene-rich, depending on the growing conditions (Tisserand and Young 2014). Western Australian Sandalwood (Santalum spicatum) is (E,E)-farnesol-rich, and Niaouli (Melaleuca quinquenervia) is (E)-nerolidol-rich (Tisserand and Young 2014). However, when normalized, these docking scores do not appear to be very significant. As such, it is unlikely that any of the examined compounds will bind viral targets (Silva et al. 2020). In contrast to other in silico studies discussed above, this study found that docking scores for carvacrol and 1,8-cineole were low. It is unclear why the binding scores were low in this study, but significant in others. Regardless, these compounds, or essential oils containing them, may warrant additional studies.
Conclusions
In silico studies can provide preliminary data to identify compounds of interest. However, in vitro and in vivo studies are needed to confirm their conclusions before implementing them as treatments for COVID-19. In silico studies have identified FDA approved medications that can be useful to treat COVID-19, and similarly, studies of naturally derived compounds may lead to treatments. Based on existing in silico studies, additional in vitro and in vivo studies examining garlic oil, cajeput oil, thymol, 1,8-cineole, carvacrol, (E)-β-farnesene, (E,E)-β-farnesene, (E,E)-farnesol, (E)-nerolidol (or essential oils containing them) are warranted.
With special thanks to Alexis St-Gelais for his input on jensenone.
Works cited
Abdelli I, Hassani F, Brikci SB, Ghalem S (2020) In silico study the inhibition of angiotensin converting enzyme 2 receptor of COVID-19 by Ammoides verticillata components harvested from Western Algeria. J Biomol Struct Dyn 1–14. https://doi.org/10.1080/07391102.2020.1763199
Astani A, Reichling J, Schnitzler P (2010) Comparative study on the antiviral activity of selected monoterpenes derived from essential oils. Phytother Res 24:673–679. https://doi.org/10.1002/ptr.2955
Boland DJ, Brophy JJ, Fookest CJR (1992) Jensenone, a ketone from Eucalyptus jensenii. Phytochemistry 31:2178–2179. https://doi.org/10.1016/0031-9422(92)80396-v
Bouchentouf S, Missoum N (2020) Identification of Compounds from Nigella Sativa as New Potential Inhibitors of 2019 Novel Coronasvirus (Covid-19): Molecular Docking Study. https://doi.org/10.26434/chemrxiv.12055716.v1
Dehelean CA, Lazureanu V, Coricovac D, et al (2020) SARS-CoV-2: Repurposed Drugs and Novel Therapeutic Approaches—Insights into Chemical Structure—Biological Activity and Toxicological Screening. J Clin Medicine 9:2084. https://doi.org/10.3390/jcm9072084
Gordon DE, Jang GM, Bouhaddou M, et al (2020) A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583:459–468. https://doi.org/10.1038/s41586-020-2286-9
Hoffmann M, Kleine-Weber H, Schroeder S, et al (2020) SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell. https://doi.org/10.1016/j.cell.2020.02.052
Jin Z, Du X, Xu Y, et al (2020) Structure of Mpro from SARS-CoV-2 and discovery of its inhibitors. Nature 582:289–293. https://doi.org/10.1038/s41586-020-2223-y
Kalathiya U, Padariya M, Mayordomo M, et al (2020) Highly Conserved Homotrimer Cavity Formed by the SARS-CoV-2 Spike Glycoprotein: A Novel Binding Site. J Clin Medicine 9:1473. https://doi.org/10.3390/jcm9051473
Li M-Y, Li L, Zhang Y, Wang X-S (2020) Expression of the SARS-CoV-2 cell receptor gene ACE2 in a wide variety of human tissues. Infect Dis Poverty 9:45. https://doi.org/10.1186/s40249-020-00662-x
Li Y, Lai Y, Wang Y, et al (2016) 1, 8-Cineol Protect Against Influenza-Virus-Induced Pneumonia in Mice. Inflammation 39:1582–1593. https://doi.org/10.1007/s10753-016-0394-3
Mollica V, Rizzo A, Massari F (2020) The pivotal role of TMPRSS2 in coronavirus disease 2019 and prostate cancer. Future Oncol 16:2029–2033. https://doi.org/10.2217/fon-2020-0571
My TTA, Loan HTP, Hai NTT, et al (2020) Evaluation of the Inhibitory Activities of COVID‐19 of Melaleuca cajuputi Oil Using Docking Simulation. Chem 5:6312–6320. https://doi.org/10.1002/slct.202000822
Qamar MT ul, Alqahtani SM, Alamri MA, Chen L-L (2020) Structural basis of SARS-CoV-2 3CLpro and anti-COVID-19 drug discovery from medicinal plants. J Pharm Analysis 10:313–319. https://doi.org/10.1016/j.jpha.2020.03.009
Sharma AD, Kaur I (2020a) Eucalyptol (1,8 cineole) from Eucalyptus Essential Oil a Potential Inhibitor of COVID 19 Corona Virus Infection by Molecular Docking Studies . https://doi.org/10.20944/preprints202003.0455.v1
Sharma AD, Kaur I (2020b) Molecular docking studies on Jensenone from eucalyptus essential oil as a potential inhibitor of COVID 19 corona virus infection. Arxiv
Silva JKR da, Figueiredo PLB, Byler KG, Setzer WN (2020) Essential Oils as Antiviral Agents, Potential of Essential Oils to Treat SARS-CoV-2 Infection: An In-Silico Investigation. Int J Mol Sci 21:3426. https://doi.org/10.3390/ijms21103426
Thuy BTP, My TTA, Hai NTT, et al (2020) Investigation into SARS-CoV-2 Resistance of Compounds in Garlic Essential Oil. Acs Omega 5:8312–8320. https://doi.org/10.1021/acsomega.0c00772
Tisserand R, Young R (2014) Essential Oil Safety: A Guide for Health Care Professionals, 2nd edn.
Tiwari V, Beer JC, Sankaranarayanan NV, et al (2020) Discovering small-molecule therapeutics against SARS-CoV-2. Drug Discov Today 25:1535–1544. https://doi.org/10.1016/j.drudis.2020.06.017
Trezza A, Iovinelli D, Santucci A, et al (2020) An integrated drug repurposing strategy for the rapid identification of potential SARS-CoV-2 viral inhibitors. Sci Rep-uk 10:13866. https://doi.org/10.1038/s41598-020-70863-9
Yadav N, Chandra H (2017) Suppression of inflammatory and infection responses in lung macrophages by eucalyptus oil and its constituent 1,8-cineole: Role of pattern recognition receptors TREM-1 and NLRP3, the MAP kinase regulator MKP-1, and NFκB. Plos One 12:e0188232. https://doi.org/10.1371/journal.pone.0188232
Yang H, Xie W, Xue X, et al (2005) Design of Wide-Spectrum Inhibitors Targeting Coronavirus Main Proteases. Plos Biol 3:e324. https://doi.org/10.1371/journal.pbio.0030324





0 Comments